The Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA) recently released updated Clinical Practice Guidelines for C. difficile Infection (CDI). The guidelines highlight several challenges facing infection preventionists, laboratory personnel, clinicians, and pharmacists regarding the diagnosis and management of CDI. This Know Your Poo Policy brief summarizes the diagnostic recommendations and focuses specifically on the role of nucleic acid amplification tests (NAATs), such as real-time polymerase chain reaction (PCR).
Summary:
- Patients with unexplained and new-onset diarrhea (≥3 unformed stools in 24 hours) who have not received laxatives in the last 48 hours are an appropriate patient population for CDI testing1
- Labs that can control the types of specimens they receive (i.e., can reject inappropriate specimens) should use a standalone NAAT or an algorithm approach that includes toxin
Testing the Right Patient Specimen
According to the guidelines, the recommended laboratory test(s) for CDI diverge based on an institution’s ability to restrict testing to the appropriate specimens collected from the appropriate patients. In order to assess appropriateness of testing a collaborative approach may be taken, which can include input from clinicians, laboratorians, infection preventionists and members of the antibiotic stewardship committee.
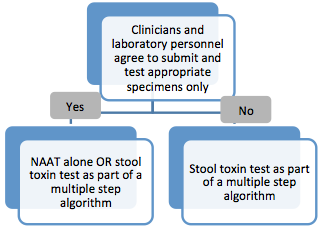
"Clinicians can improve laboratory test relevance by only testing patients likely to have C. difficile disease. This includes not routinely performing testing on stool from a patient who has received a laxative within the previous 48 hours. Laboratories can improve specificity by rejecting specimens that are not liquid or soft."1
— IDSA Guidelines
|
Click here to access complimentary Clinician Education Tools |
Test Sensitivity
While the new guidelines mention several approaches for diagnosing CDI, including those that utilize multi-test algorithms, PCR remains a highly sensitive method for identifying toxigenic C. difficile, which is critical for both accurate diagnosis and infection prevention activities in hospitals.
The guidelines note the importance of utilizing a highly sensitive test method and cite studies in which PCR was found to be more sensitive than other test methods. The first, by Peterson et al. found that PCR (93%) was more sensitive than toxin EIA (73%) and direct cytotoxin testing (77%).1 "The authors concluded that PCR outperformed the other diagnostic test methods when applied to patients who meet clinical criteria for C. difficile disease."1 A second study, Berry et al, assessed whether a rapid PCR assay (GeneXpert C. difficile assay) or a GDH-toxin algorithm correlated well with CDI diagnosis. They found that the GDH screen did not detect 16% of patients with clinical CDI. In addition, when the GDH screen was combined with a toxin test (GDH-algorithm) 60% of patients with clinical CDI would have been missed.2
1. Clifford McDonald, et al. Clinical Practice Guidelines for Clostridium difficile Infection in Adults and Children: 2017 Update by the Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA). Clin Infect Dis. 2018 Feb 15. [Epub ahead of print]. Accessed Mar 2018. https://doi.org/10.1093/cid/cix1085
2. Peterson LR, et al. Detection of toxigenic Clostridium difficile in stool samples by real-time polymerase chain reaction for the diagnosis of C. difficile-associated diarrhea. Clin Infect Dis. 2007 Nov 1;45(9):1152-60.
3. Berry N, et al. Real-time polymerase chain reaction correlates well with clinical diagnosis of Clostridium difficile infection. J Hosp Infect. 2014 Jun;87(2):109-14.
 .
.